Now that we’re getting closer to analyzing the atmospheres of terrestrial-size exoplanets, it’s worth remembering how difficult the call on the existence of life is going to be. Long-time Centauri Dreams contributor Alex Tolley takes on the issue in his essay for today, pointing out along the way just how easy it is to see what we want to see in our data. While we can learn much from terrestrial biology, new approaches looking at ‘pathway complexity’ may offer useful indications of biology and a set of markers not constrained by our own unique sample of life on Earth. A lecturer in biology at the University of California, Alex brings us up to speed with extending our methods of life detection in ways that are ‘biology agnostic.’ Expect controversy ahead — will we know life when we see it, and how can we be sure?
by Alex Tolley

Manuel Werner, CC BY-SA 2.5, https://commons.wikimedia.org/w/index.php?curid=633977
Life: [noun]? The condition that distinguishes animals and plants from inorganic matter, including the capacity for growth, reproduction, functional activity, and continual change preceding death. – Oxford Living Dictionary [6]
Life, like pornography, is notoriously hard to define, but we mostly recognize it when we see it. Life, as we know it, is identified by a set of features, which individually, may be shown by non-living systems. A classic example is “fire”, that can exhibit a simple metabolism (combustion), growth (size and spread), and even “reproduction” (sparks ignite new fires). Fire, however, fails the test of life, as all terrestrial life has the cell as a basic unit, which is not a feature of fire, nor can fire evolve.
Fossils are clearly not living, yet they show the order that life exhibits which indicates that they are a remnant of an organism that was living. For example, the fossilized skull of a dinosaur shows considerable order with features that indicate it was from a living animal and very similar to other fossil skulls of its type. Fossil bone fragments are far harder to identify and experts can detect these when a layperson would see only a piece of rock. Microfossils are even harder, as the controversial objects in the meteorite ALH84001 indicate [7]. Are they natural formations or organisms?
When we consider how to recognize extraterrestrial life, we are largely constrained by the single sample we have. That should not stop us looking for Earth-type life as the low hanging fruit, as Earth-type life is an existence proof and well worth searching for signs of, whether with telescopes or probes.
Recent focus has shifted to spectroscopic analysis of exoplanet atmospheres. The logic is largely that of James Lovelock’s Gaia hypothesis, where the production of certain gases is a proxy for their generation by life. For a terrestrial-like world in the habitable zone (HZ), the existence of both oxygen (O2) and methane (CH4) implies life as these are primarily produced by life on Earth in the ratios required to prevent equilibrium. For a world more like the Archaean Era, an atmosphere rich in methane but excluding other gases like carbon monoxide (CO) indicates bacterial methanogens, as the geological serpentinization of ultramafic rocks like olivine is insufficient to maintain the CH4 levels.
It is this geological reaction that makes the presence of CH4 in the Martian atmosphere so ambiguous, as the masses are small enough to be produced by geology as well as subsurface pockets of life.
Where we have extraterrestrial samples, such as carbonaceous meteorites, asteroids and the recent confirmation of organic material on Mars, there is a need to differentiate abiotic from biotic processes. The classic examples of biotic processes based on our Earthly sample include the chirality of amino acids and sugars, the isotopic changes of elements due to favored selection in biological processes, such as the reduced carbon-13/carbon-12 ratios, and the odd number of carbon atoms in many lipids.
As our planetary probes increase in sophistication, and the idea that subsurface icy moons might be hospitable for life, there is a need to include the instruments to test for possible biosignatures to try to reduce ambiguity.
Biology as a System
Returning to the question of recognizing life, a key point is that it exhibits a number of features that need to be present so that we can distinguish it from inanimate objects. For terrestrial life, the basic unit is the cell, which encapsulates all the components needed to exhibit the features we identify with life, maintaining order and fighting entropy, by interacting with the external world. With the evolution of photosynthesis, that order is maintained by the capture of a tiny amount of the energy emitted by our sun. This is now the dominant source of energy for the terrestrial biosphere. Even the simplest unicellular organisms require hundred of genes, and therefore unique functional protein molecules to maintain themselves. Higher organisms require tens of thousands of genes, producing hundreds of thousands of unique proteins to maintain their more complex structures and life cycles.
We can again see the problem of detecting life from limited features with the three Viking experiments that proved ambiguous. Had there been a microscope to view a culture, the presence of cells, their growth and reproduction over time would have clinched the presence of life.
While this approach can work for samples in our solar system, for exoplanets, we must rely on proxies that are primarily measurable using spectrographic techniques. Conceivably, a telescope could image a world, detecting seasonal changes in photosynthetic organisms, providing direct evidence. For worlds with life still only in its prokaryotic state, remote direct imaging of life may prove impossible.
For samples in our solar system, we can expect a search based on terrestrial life analogs, so the usual suspects will be searched – proteins with chiral amino acids, DNA, lipids with odd-numbered carbon atoms, as well as more subtle signs such as carbon-13/carbon-12 ratios. But we should also look for evidence that is terrestrial biology agnostic, especially if we are hoping to discover very different life forms from unique geneses.
Sara Seager: Going Beyond the Presence of a Molecule
Sara Seager’s team has been at the forefront of considering biosignatures beyond the usual proxies of atmospheric gas mixing ratios. Her paper [8] (see also CD post Ambiguity in Life Detection?, October 31, 2017) collated the range of small organic molecules that exist and their source whether biotic or abiotic or both. At the 2018 Breakthrough Discuss conference, she noted that biology does have some apparent constraints and explained the paucity of biotic molecules with nitrogen-sulfur (N-S) bonds, even though these compounds abound in industrial chemistry because of their usefulness. Terrestrial biology is rich with thiol reactions and has evolved replication and metabolisms that generally eschew molecules with such N-S bonds. This phenomenon constitutes a possible biosignature. While this is one specific example, there are likely many others. However, constraining our ideas to terrestrial biology may result in us missing non-terrestrial biologies that are different, providing false negatives. What is needed is a more general approach that is biology agnostic.
Lee Cronin: A Generalized Approach
Lee Cronin’s group has been formulating a more biology agnostic approach, one that is based on living organisms being homeostatic systems [4]. His approach is to assume molecules can be constructed by assembling sub-units and that this confers a minimal construction pathway, which he calls “pathway complexity”. One can consider this as a tree of all possible molecules composed of the building blocks with the number of construction steps needed to build the molecule. This is an indication of the non-randomness of the molecule. If a molecule in a sample is highly enriched compared to the possible random set of molecules that could be constructed at random, this is indicative of a construction pathway that in turn is indicative of life.
Figure 1 below shows the concept of construction of a specific molecule from building blocks.
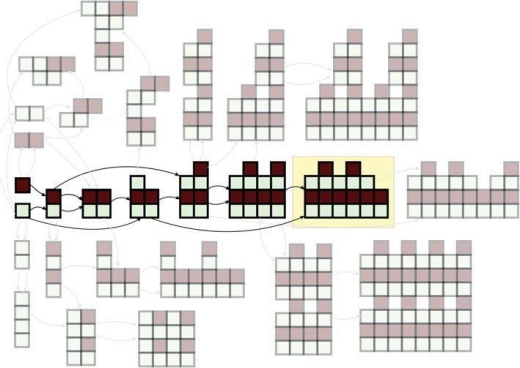
Figure 1. Illustration of a complexity pathway in blocks, with the target shown by the yellow box. A combinatorial explosion in structures is illustrated by the other faded structures shown, which are just a small set of the many alternative structures that could be constructed. (Online version in color.) [4]
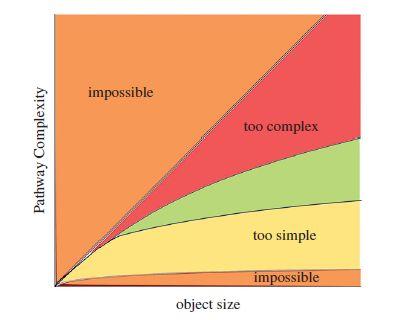
Figure 2. An illustrative graph of complexity against size of the state space. Orange regions are impossible as they are above or below the bounds of the measure. The green region is where living systems may be most probable, where structures are neither too simple to be definitively biological, nor too complex to exist at all. [4]
Cronin states? :
“We can extend the basic complexity measure above to cope with assessing the complexity of a group of objects that contain identical connection motifs (figure 5). In this case, we examine a population of objects and abstract out a common graph based on connected subunits that share features. For example, if examining a set of cups or mugs, then we can create a common graph of ‘handle connected to body’, regardless of potential variations in size/colour etc. If examining a set of human beings, then we could create a common graph of bone connectivity, ignoring variations in size/shape of individual bones, or any material in the body other than bones.”
He concludes:
“It is clear that biological and biologically derived systems have an ability to create complex structures, whether proteins or iPhones, that is not found elsewhere in nature. Assessing the complexity of such artefacts will be instrumental in searching for undiscovered biospheres, either on Earth [29] or elsewhere in the Solar System, and would make no assumptions about the details of the biology found. We propose Pathway Complexity as the natural measure of complexity for the production of artefacts. In this context, we argue that there is a critical value of Pathway Complexity above which all artefacts must be biologically derived. This approach provides a probabilistic context to extending the physical basis for life detection proposed by Lovelock [30]. In further work, we will show how this applies to a range of other systems, and propose a series of experimental approaches to the detection of objects and data that could be investigated as a possible biosignature. In the laboratory, we are interested in using this approach to develop a system that can explore the threshold between a non-living and living system. Pathway Complexity may also allow us to develop a new theory for biology. This might inform anew way to search for life in the laboratory in terms of the complex products a system produces and if they could have arisen in any abundance by chance, rather than trying to measure the intrinsic complexity of the living system itself.”
Kauffman: Self Organization Theory of Life
In 1993, Stuart Kauffman published The Origins of Order: Self Organization and Selection in Evolution [2]. I consider it a tour de force? in theoretical biology. Of relevance to this post are chapters 7 and 8 on the concept of autocatalytic sets, and the crystallization of metabolisms.
Autocatalytic sets are best thought of as a linked set of components, e,g, catalytic RNA, that can build each component from others in the set. The effect of which is to rapidly increase the RNA species included in the set. From an origin of life perspective, Kauffman showed that the probability of autocatalytic sets arising increases to unity as the number of RNA species increases. Metabolisms similarly crystalize when there are enough reactants so that a complete, self-contained metabolism can be sustained. Again, the probability of such a complete metabolism will increase as the number of reactants increase.
As Kaufmann states:
“Thus we arrive at a new point of view. The emergence of a connected metabolism as a supracritical web requires a sufficient complexity of organic molecules and a sufficient complexity of potential catalysts. At that point, such a connected web is an inevitable emergent collective property of the chemical system.”? [2] p348.
The relevance to Cronin’s work should be clear. Once a self-sustaining set of components appears, the components in that set will increase rapidly compared to others in the vast space of possible components. Cronin’s metrics, such as “pathway complexity” naturally emerge when considering the number of components compared to the possible components due to random reactions.
While Kauffman’s work is theoretical, Cronin has shown that lab experiments [5] support this basic concept. In terms of biology, they are agnostic in origin, therefore freeing us from focusing on terrestrial biology as our single sample of life, and informing us of possible biosignatures.
Biosignature Search
Sampling the compound space is not something that is likely to be possible anytime soon, if ever, using spectral analysis of [exo]planet atmospheres. Even Seager’s list of possible biosignature compounds are effectively trace compounds, and there is no way to determine whether her N-S bonds hypothesis works for an exoplanet from telescopic observations.
However, the solar system is another matter, and targets such as Mars, the plumes of Enceladus, the organics on the [sub]surface of comets, Ceres and the Europan sub-surface ocean are ripe for this sort of systems analysis using mass spectrometers and IR spectroscopy on probes to determine the mix of compounds in a physical sample.
While future telescopic observations that can image worlds directly may show up life as lush, boreal zones on exoplanets, nearer to home we may be able to sample the biological detritus of such worlds through wanderers like ‘Oumuamua that may have captured bacterial life from living worlds. If exoplanet life is largely bacterial, then probes sampling the upper atmosphere or even the surface can use this technique to determine if life exists without the difficulty of trying to cultivate bacterial colonies and observing the results. While interstellar probes that could sample such worlds are a relatively distant prospect, they are possible in the centuries to come using propulsion technologies that do not require new physics.
Conclusion
Confirmation bias involves seeing the data supporting what you are expecting. The lack of artificial objects in the heavens that is the context for the Fermi Question elicits polar views of “we are the only life” to “technological life is there, we don’t recognize it”. Similarly, the lack of unambiguous signals found by SETI results in a similar dichotomy. As we noted, the ambiguous objects in the ALH84001 meteorite that came from Mars have proponents for either proposition — life and non-life. Early searches for “missing links” in the evolution of humans that found a few bones and partial skulls also resulted in polar views of whether modern humans had evolved from an apelike ancestor or been created in his present form. We can be sure that any spectroscopic evidence of proxies for life – biosignatures – will be similarly interpreted.
So far, chemical analysis of samples in our solar system have been teasers, hinting at possible life, but no more. While a video of a living animal in a sample tube would be unambiguous (although there will no doubt be claims of “it is a hoax”), the most compelling approaches would be confirmation of DNA or proteins, preferably with no known terrestrial copies. This, however, assumes life is very similar to terrestrial life, and techniques used to find such molecules will miss life that may be very different from terrestrial life. Starting from the model that living systems are complex systems, yet not so complex as to be random, chemical analyses within the scope of that with existing analyzers may well be able to indicate life with far less ambiguity than the focus upon a few proxy molecules. In this regard, the theoretical bases described by Kaufmann and Cronin, and confirmed with experiments on terrestrial living organisms, offers perhaps the best approach for sampling probes that we can envisage in the near future, although the mass penalty of a microscope would be very much appreciated.
References
1. Petkowski J et al “Natural Products Containing a Nitrogen?Sulfur Bond” J. Nat. Prod. 2018, 81, 423?446
2. Kauffman S. “The Origin of a Connected Metabolism” ch 10, p343 in The Origins of Order, 1993
3. Domagal-Goldman S et al “Life Beyond the Solar System: Remotely Detectable Biosignatures” 2018, arXiv:1801.06714 [astro-ph.EP]
4. Cronin Lee “A probabilistic framework for identifying biosignatures using Pathway Complexity” 2017, Philos Trans A Math Phys Eng Sci. 2017 Dec 28;375(2109). pii: 20160342. doi: 10.1098/rsta.2016.0342.
5. Doran D et al “A recursive microfluidic platform to explore
the emergence of chemical evolution” 2017, ? Beilstein J Org Chem.? 2017 Aug 17;13:1702-1709. doi: 10.3762/bjoc.13.164. eCollection 2017.
6. http://en.oxforddictionaries.com/definition/life
7. http://en.wikipedia.org/wiki/Allan_Hills_84001
8. Seager, Bains and Petkowski, “Toward a List of Molecules as Potential Biosignature Gases for the Search for Life on Exoplanets and Applications to Terrestrial Biochemistry,” ? Astrobiology 16(6) (June 201), 465-485



A test for growth of single celled life such as a series of small sealed vessels containing simple carbon, hydrogen, oxygen, nitrogen, sulphur and phosphorous containing compounds in an aqueous medium and heated to some temperature between 20C and 50C with various gases present might produce growth. If this test module is linked to a series of detectors for changes in optical density, emissions of gases etc. we should at least be able to determine whether bacteria-like organisms are present on Mars or other solar system bodies. Sampling beneath the highly oxidizing surface is obviously a key part of a Mars survey. The Viking landers collected soil samples and subjected them to tests via a GCMS. This has clear difficulties as was concluded at the time. Surely the type of experiments I have described are possible now in small sealed containers? I’m surprised some version of this hasn’t already been tried. Sample recovery from Mars for return to earth is surely much more technically difficult and fraught with danger for any biological organism contained within the sample.
This seems like an obvious approach and indeed was one of the Viking experiments. However, even on Earth, very bacteria can be cultured at all. Culture medium exposed to organisms usually just capture just a few bacteria species. Sequencing all DNA in samples, es Venter did for ocean samples around the world shows that there are vastly more microorganism species in the wild. Arguably, we don’t care, as long as just one or a few exotic life forms will replicate in artificial media. If however, they are like most bacteria, they just will not replicate and the experiment will fail. We won’t know if the lack of growth is a true or false negative. A lab on Mars might well spend some effort to try to culture swabs of subsurface regolith, ice, and water this way. For a mass limited space probes, sampling the plumes of Enceladus, or a few scrapings from an asteroid or comet, this might not be the most efficient test for life, especially if it eventually proves quite “exotic”. The Viking experience should be a cautionary example, especially as we have very different technology to detect life today.
I don’t think we can use Mars as an example for life since there has not yet been a detection of life there. There may be fossilized evidence of microscopic life there. I don’t understand the idea behind this paper. It seems to me absurd to look for ET life which is different from Earths and has a different biosignature since we don’t know what to look for and there is no evidence for the physical existence of such life. It’s better to look for what we know. Gnosis means knowledge of the spiritual and agnostic means not knowing or we can’t know anything beyond the physical phenomenon. I don’t get the use of the term “biology agnostic.” There is no such thing as a non agnostic biology as far as science is concerned.
If it walks like a duck, quacks like a duck and tastes like a duck, could it have been copied from the real thing by a von Neumann probe?
Broadening the scope of a search for life is not absurd. It is important to remain rational but also open to options that may seem unlikely yet have a sound theoretical basis. Your suggestion that we search primarily for life in the form we are familiar with is already the case and will continue to be so. Fortunately we can also look for other options at the same time.
The point I was trying to make is that while we should certainly search for “life as we know it”, we need less ambiguous signatures for that life. As life may be somewhat different from terrestrial life, we should also aim to detect life using phenomena that is generally indicative of life, rather than searching just for what we know.
Consider for a moment the idea broached many times in CD articles that pre-photosynthetic, prokaryotic life is likely the most common life on exoplanets. Using the terrestrial analogy, we, therefore, look for worlds with strong CH4 signals in a mainly N2-CO2 atmosphere, because we expect that life to be methanogenic. But what is our assumption is wrong, and life on many worlds has a different biology, or CH4 is not a metabolite? A more general approach is to look for any gas mixtures that are out of equilibrium.
For our solar system, we will probably want to use a battery of tests to search for life, some of which assume terrestrial biology, others that extend the tests to more exotic life.
One final point, we call studies of life-like software as “artificial life”. As Robin Datta suggests, what if there are artificial life forms that mimic life. They don’t have to be large von Neumann replicating probes, but potentially microscopic. A finding of such artificial life would have tremendous implications, as long as we can recognize it. A bit more “down to earth”, Paul Davies suggested looking for a “shadow biosphere” on Earth to see if there is “life as we don’t know it” here on Earth. It might be difficult to search for, but it isn’t a crazy idea.
A very interesting article.
I have been of the opinion for some years now that the best use for SETI funding is new propulsion/spacecraft development instead of searches by telescope. We must go out there if we want to find life.
By definition, SETI funding is for searching for extraterrestrial intelligence. The only technologies we have at present for this are remote viewing of electromagnetic radiation by telescopes. At this stage, new space travel development would be a separate funding budget. In the US, this funding is mostly NASA’s NASA Innovative Advanced Concepts (NIAC).
Extraterrestrial Intelligence can be biological, variant biological (but still carbon-based), non-carbon based biological, and post-biological, based on initial uploading of intelligence to systems based on technologies along the lines of electronics.
Biosignature atmospheric compounds that cannot exist in the absence of intelligence, and megastructures including Dyson spheres, ONeill cylinders, etc. are all things to look for in existing and future datasets, where even a single instance of irrefutable evidence may clinch the answer to big questions.
I agree that directly exploring other star systems and regions of the galaxy which seem like good places for alien life would be our best bet to find and confirm their existence.
However, since anyone who reads Centauri Dreams even casually knows that real interstellar travel of the kind that will only take decades instead of centuries or more is still far away, even the much vaunted Breakthrough Interstellar project, we will have to rely on radio and optical telescopes for a long while to conduct SETI and search for biosignatures. It is far better than nothing and gives us a chance to put some solid data to our speculations and theories, even if that data is a negative (we can know where the really noisy technological aliens are not, for example).
Astronomers have done some pretty clever sleuthing despite being many light years from their subjects of examination. This especially applies to now but even centuries ago. This certainly applies to SETI and Bioastronomy.
Excellent article, thanks Alex!
Two instruments are most useful here:
1) a 2d chromatograph/mass spec device that can image molecular complexity
2) a microscope to image spatial complexity (as Alex has also called for)
Hopefully, NASA will start including more life-detection instruments in future missions. I understand they dialed that back primarily because of the problems with the Vikings
That was good, Alex Tolley!
I am tending towards seeing it all as a continuum, from (pre-chemical) particles through inorganic, organic and bio chemistry and on to molecular cell biology, multicellularity, division of labor, intra- and inter- species interactions with sociality and ecosystems.
“Life” is a certain set of phenomena that, with the proper starting conditions make their appearance and evolve, and in so doing, creating their own milieu.
What is the range of the “proper starting conditions” and how long would it take are among the questions that arise.
Can survival, growth, replication and evolution / learning be so subtle or advanced that it will be overlooked by humans?
Might we miss manifestations of sentience? If we acknowledge Paul Stamnets, mycelia might be categorized as sentient right here on earth.
There may be more leeway with Fermi and Drake than we are given to realize.
Variously attributed:
“Not only is the Universe stranger than we think, it is stranger than we can think.” ? Werner Heisenberg
Not only is the universe stranger than we imagine, it is stranger than we can imagine.
Though sometimes attributed to Arthur Eddington without citation, this seems to be derived from a statement by J. B. S. Haldane, in Possible Worlds and Other Papers (1927), p. 286: The Universe is not only queerer than we suppose, but queerer than we can suppose.
Thank you so much, Paul, and especially Alex, for this terrific rundown that even this non-scientist has a broad overview. It is much appreciated.
What exciting times we live in! Not only do we not know if there is life outside Earth, we don’t even know what life IS. Truly this is a problem with scientific as well as philosophical components.
The scientific and ethical progress made by human kind indeed contains much to marvel. In reality, though, we know next to nothing about this curious place that we find ourselves, the place we call the “Universe”. So many questions remain- the nature of non-baryonic matter being in my view at the top of the list as surely the answer will contain the key to so many related, corollary questions.
It’s my own prejudice, one that perhaps Mr. Tolley would question, as he ponders the observed ability of matter to organize itself into Life.
Whatever that is.
In my view a light microscope is of limited usefulness as we are limited to the wavelengths of visible light, so 1000x at best with oil immersion. And a very compact electron microscope is not yet available is it? Looking for small bacteria will remain a problem by microscopy and the Mars microbes may be very small if indeed the ALH 84001 possible “microbes” are anything to go by. The same applies with virus-like entities. I’m thinking we’re going to find microscopic life in the solar system first, then possibly more exciting things out in other stellar systems. So we have to develop a suite of methods for detecting microscopic life on other planets and moons within our solar system.
If it moves (sometimes not always), consumes, produces waste and reproduces we should be able to detect it. Surely at least for microscopic life. That we haven’t put the right types of probes on the ground on Mars at the very least shows a lack of endeavor and imagination. We roll around on the surface looking at rocks and do very little else or so it seems. I realize we have mapped the planet very well and are very sure now that Mars was once a wet planet but the life sciences are lagging woefully behind. A few feet below the surface in appropriate locations may lie huge numbers of microorganisms. Do we know much about the methane flux on Mars and possible source regions? So much to do and so little has been done so far.
Yeah, we need to send astronauts there. Rovers are simply too primitive for the job.
I like the general thesis Alex but let’s get going on the more practical, pragmatic one step at a time approach. If we can’t dig down a few feet on our nearest planetary neighbor after decades of being able to land probes there what are we doing? If we have to go there ourselves to set up a sophisticated enough lab to test properly for microscopic life then let’s get going! No wonder Robert Zubrin is losing his hair (no offense intended Robert). I’m pulling mine out with frustration as well. Can we not put together an international consortium as with the space station and get to Mars in the next 10 years or so?
I agree. When Beagle 2 crashed on Mars why didn’t various stakeholders join forces to send Beagle 3? The Americans have gone so slowly and gone so far to exclude life search after failure of Vikings to confirm/deny life. If you are not failing often then you are not trying hard enough.
I wonder if any Petrie-dish type experiments have been done on retrieved samples from space? I have never heard of these being done although it would seem an obvious thing to try. Hope it is not a case of: “For safety reasons all samples must be sterilised prior to testing.”
It seems obvious that we haven’t tried anywhere near hard enough to detect life on Mars Coacervate. We’ve waited decades since the Viking landers. What is going on? It seems to obvious that it should be tried in earnest again. We have improved the technology and hardware enormously over the intervening time and yet each probe gets launched with the proviso “we won’t be able to determine with any certainty whether life exists on Mars with this probe.” Wasn’t the Mars Science Lab the perfect opportunity to include a decent sized drill and a life detection platform of some kind? I’m very puzzled about this. Have we become terrified of failure?
NASA went to Mars with Viking 1 and 2 under the presumption that microbial life would be found there on the surface by “merely” scooping up some soil and dumping it into the twin landers’ $60 million (in 1975 dollars) compact automated biology labs.
This was done with a very incomplete picture of what the Martian surface was composed of and a lingering hope from the days of Percival Lowell that while Mars may not have been as lively now as it was once presumed to be, there might still be some hardy creatures managing to survive and reproduce on the otherwise harsh planet. We still hold out such hope even today, but we now think it is well under the Martian surface.
Despite the amazing success of both landers and their orbiters in otherwise revealing Mars, the fact that the Vikings did not return unambiguous evidence of native organisms made NASA gun-shy about searching for life there for decades. That is why the rovers Spirit and Opportunity were only designed to look for signs of past water (they could have also searched for fossils and may have found some, but watch NASA mission scientists scatter at that word). Even Curiosity, which had been originally touted as an actual Martian life searcher, was downgraded to finding the organic chemicals that make up life. As for finding any fossils – Yikes!
Yes, we need a bolder and wider effort to explore our Sol system for life. We know there are numerous places besides Mars which frankly offer much better chances for supporting life. This even includes Venus both high in its atmosphere and under its surface!
Oh yes – remember how long it took for mission scientists to confirm that the droplets clinging to the metal legs of Mars Phoenix in 2008 were actually liquid water? A child could tell you that was moisture, but even a few water drops were frightening to confirm at first.
It is one thing to be scientifically conservative, but the agency was more concerned about prematurely declaring we are not alone and looking foolish. We are much too cautious society these days as it is.
And then there was the Wolfe-Simon arsenic life debacle when Nasa got ahead of itself.
Elon is our only hope.
Or Jeff Besos. Or China. Or maybe India and Russia.
I’ve been complaining about this for a long time now. NASA has been taking direction from geologists and planetary science folks, not from experimental biologists, and this has to change.
Quote by Alex Tolley: “Consider for a moment the idea broached many times in CD articles that pre-photosynthetic, prokaryotic life is likely the most common life on exoplanets. Using the terrestrial analogy, we, therefore, look for worlds with strong CH4 signals in a mainly N2-CO2 atmosphere, because we expect that life to be methanogenic. But what is our assumption is wrong, and life on many worlds has a different biology, or CH4 is not a metabolite? A more general approach is to look for any gas mixtures that are out of equilibrium.”
Biologists and scientists already know that the potential ubiquity of Carbon based life is not an assumption and nature’s choice of it is not random, but based carbon being the most versatile of elements compared to other elements. Nature is based on efficiency and strength. Carbon has four valence electrons, so it can give up four of them or receive four electrons to complete an electron shell. It can combine with many other elements to form compounds and form long carbon chains.
There is a lot more space in carbon chemistry before we consider unlikely silicon or other element chemistry. Every aspect of life’s functions may use very different molecules than those on Earth: information storage, structural and catalytic proteins, metabolites…. When we have a large sample of exo-life from other worlds, much like Kepler has given us of exoplanets, we may find life ranges from very similar to Earth life, varying in few aspects, to very different. Imagine life that evolutionary converges with Earth’s, but with a different genetic code (we are close to modifying life with a new genetic code today. We already have developed translation with 4 base codons and novel amino acids).
Thanks for your response Alex. I didn’t notice until now. I agree we haven’t been very successful getting microbes to grow in standard culture media. The idea of not trying to extend our knowledge seems unsatisfying though. If something is hardy enough to grow under the soil on Mars maybe it will be easier than we think to get at least some response in terms of reproduction and generation of waste products, i.e. altering the composition of the growth medium. We’ve waited 42 years since the Viking landers to try hard to detect signs of life. Isn’t that a long enough interval? Waiting longer won’t make it any easier. Surely we can’t be that bereft of ideas that we can’t even try?
You might find this link about detecting bacteria in the Atacama desert as a Mars analog interesting.
Life at the dry edge: Microorganisms of the Atacama Desert.
Of the many and varied soil microbes, bacteria in numbers of individual cells can be a billion to a gram of soil.
https://en.m.wikipedia.org/wiki/Soil_microbiology
DNA analysis indicates a million species of bacteria in 30 grams of soil, around 1% of which can be grown in the lab.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3160642/?report=classic
The failure to grow most bacteria may be due to multiple dependencies, e.g. the growth of A requires the growth of B but after C has grown and stopped growing; C requires the growth of D but without growth in E, etc.
The main problem to grow bacteria is the culture media. But there is a solution:
https://www.popsci.com/ichip-new-way-find-antibiotics-and-other-key-drugs#page-3
Biospheres off earth, then, will need to be huge enough to have energy exchanges between components due to mass and be able to transform cosmic radiation into useful energy to escape solar systems.
There must be an upper limit to what machinery itself can maintain — the interior of mountain sized asteroids, for example.
More difficult a task than many imagine.
Great article, thank you.
I think it makes convergent evolution easier to understand. If RNA speciation must continue, then the occurrence of any gene becomes more likely, even inevitable. Being 1 out of 50 billion is still exceptional, but also inevitable, if the catalytic process can continue adding 1 to the count.
Kauffman’s theory makes a compelling case that ambiogenesis is common. Does it also imply that cell membranes are a required first step for ambiogeneis? If so, there are many lipid types that could form membranes and geological processes to catalyze membranes, causing membranes to speciate. It may also make protolife common, stalled steps on the way to a membrane becoming autocatalytic.
I hope we find life to be nothing less complicated than the soapy lubricant celestial calalytic machines refine.
Thanks for the reference Alex. Yes, I follow the work of Chris McKay quite avidly. The Atacama is a very useful model. Some of the more technical aspects of various types of “omics” would be difficult to use in a Mars setting. PCR and other detection strategies for low numbers of DNA (or RNA) containing microorganisms require primers which in turn require some knowledge of the sequence of DNA or RNA in the organism being studied. This would be very difficult to predict for an alien species. Other methods require quite large instruments (such as GC Mass Spec). We have our work cut out for us looking for alien microbes elsewhere in the solar system. I look forward to seeing novel strategies for “alien hunting” in the near future.
You are thinking that primers are needed for replicating the samples? DNA sequencing is advancing such that this is not always necessary. If Martian life is biologically the same as Earths, then chips with all combinations of complementary oligotide sequences can be used to detect, and ultimately analyze DNA and RNA sequences. But yes, this does assume some knowledge of Martian life, which may be unjustified if the underlying assumptions are wrong. The more exotic life is, the less useful some of our molecular biology techniques become, pushing us back to earlier, simpler methods, like direct observation if possible. But once detected, the vistas of new biology that would open up, with the full gamut of adapted techniques from terrestrial biology being developed and used. The journal literature would just explode.
DNA, RNA and protein coats of viruses have carbon in them. Any life found on Mars would have to be just like Earths, four base pairs, etc.
Amino acids beyond the standard twenty, nucleotdies beyond the standard five and expanded genetic codes have all been functionally incorporated into biologic systems in the lab.
https://en.m.wikipedia.org/wiki/Xenobiology
Even so much as a variant chirality could vitiate a common descent.
“Xenobiology”. I must definitely add that word to my lexicon in this context. Most of the links in that Wikipedia article are what I would consider synthetic biology, but I like the more evocative term when discussing alien biologies that we might find.
You will then also enjoy the term – and the following literature – Xenology:
http://www.xenology.info/Xeno.htm
What is your logic behind this statement? For example, why not 2 base pairs, or 6?
I have seen information theory approaches to this question that propose 4 base pairs as the optimal number for a self propagating system. 2 base pairs would be too stable making mutations unlikely. 6 base pairs would be too unstable, mutations would be too common. I will try to track down some papers.
http://www.math.unl.edu/~bdeng1/Papers/DengDNAreplication.pdf
Abstract
“In this paper we construct a mathematical model for DNA replication
based on Shannon’s mathematical theory for communication. We treat DNA replication as a communication channel. We show that the mean replication rate is maximal with four nucleotide bases under the primary assumption that the pairing time of the G–C bases is between 1.65 and 3 times the pairing time of the A–T bases.”
Please forgive the multiple posts. The paper above doesn’t address mutation rates. It proposes that 4 base pairs provide an optimal data transmission rate. 2 base pairs replicate faster but propagate less information. 6 base pairs propagate more information but at a slower speed.
This paper looks at just one aspect of DNA – base pairing rate. There are so many other factors that probably contribute to replication advantage, even during the pre-biotic stage when RNA/DNA appeared and locked in their structures. While base pairing will certainly help in replication, whether this is the key parameter to success is not proved by this paper. As the authors note, a 2 base system (G-C) may have preceded the 4 base system. 4 bases and a 3 -base codon are very good at robust coding for the amino acids that are used in proteins. However, we are still unclear how these 2 systems co-evolved. While the RNA world first model would support the base pairing rate argument, so far chemists have failed to show how this RNA world can start, although there are some intere4sting ideas. The metabolism first model would suggest that protein catalysts are more important and that coding increasingly complex proteins would drive the evolution of RNA/DNA.
Even if the 4 base model is decisive on Earth, it does not mean that alien biologies will be using the same bases for RNA and DNA. The synthetic biology experiments to add new bases and amino acids show that there are other possibilities.
Acquiring life samples that started with a separate geneses from that on Earth will provide empirical answers to our speculations.
I agree, though I would suggest base pairing rates would always be a primary factor in determining how many base pairs an ecosystem employs. An ecosystem is likely to select for the highest data transmission rate possible.
https://en.m.wikipedia.org/wiki/Nucleobase
“Five nucleobases—adenine (A), cytosine (C), guanine (G), thymine (T), and uracil (U)—are called primary or canonical. They function as the fundamental units of the genetic code, with the bases A, G, C, and T being found in DNA while A, G, C, and U are found in RNA. Thymine and uracil are identical excepting that T includes a methyl group that U lacks.”
https://en.m.wikipedia.org/wiki/Artificial_gene_synthesis#DNA_synthesis_and_synthetic_biology
“In 2012, a group of American scientists led by Floyd Romesberg, a chemical biologist at the Scripps Research Institute in San Diego, California, published that his team designed an unnatural base pair (UBP).”
“This is the first known example of a living organism passing along an expanded genetic code to subsequent generations.”
“The successful incorporation of a third base pair is a significant breakthrough toward the goal of greatly expanding the number of amino acids which can be encoded by DNA, from the existing 20 amino acids to a theoretically possible 172, thereby expanding the potential for living organisms to produce novel proteins.”
We should be cracking on with exobiology on Mars. It’s far past time to begin in earnest. Surely a broader approach would be very worthwhile. A first attempt at realistic experiments before we make efforts in much more difficult environments such as Ganymede. This has been bothering me for quite some time.
TESS has recorded OVER ONE THOUSAND transit-like events in JUST ITS FIRST DOWNLOAD!!!!! How many of them will be planets? How many of THOSE will be potentially habitable? How many of THOSE(SPECIFICALLY ~1.7Re “LHS 1140b” types will we have the ability to detect life on, should there be any life to detect?
This video link explains the TESS mission and the exquisite deployment to its orbit. Given its nominal 2 year mission, it will be interesting to see what it discovers in that period, with data downloads every 2 weeks. [ Harry’s caps lock key will wear out. ;) ]
How about the cheela – sentient beings on the surface of a neutron star (not a planet) in Robert Forward’s Dragon’s Egg?
https://en.m.wikipedia.org/wiki/Dragon%27s_Egg
When will we have the means to detect them?
While the concept of intelligent life (or any kind of life) on a neutron star is fascinating, their portrayal as characters was less than imaginative in my opinion.
Despite looking like tiny slugs, the Cheela had a history, culture, and general behavior that resembled humanity perhaps a bit too much for me to accept considering their environment and what it did to their physiology. Shades of Star Trek and so many other science fiction stories that have a great and unusual idea but then neglect the rest such as alien biology and thinking.
But hey, let’s send an expedition to some neutron stars so we can discover if they do have life and how they act.
A whole slew of papers regarding biosignatures is now up on exoplanet.eu. Way too many for me to report on here. ljk et al: Check them out and post the best ones here, please.
Thank you, Harry. I will start by making a link to their Bibliography page and letting folks peruse it:
http://exoplanet.eu/bibliography/
Alex Tolley: A must read! On the NASA Astrobiology Institute website(https://nai.nasa.gov)right now: “The momentous transition to multicellular life is not so hard after all.” Re: An article in the journal, Science, authored by Matt Herron and Will Ratcliff.
A fading Martian Opportunity
NASA’s Mars rover Opportunity has been out of contact with the Earth for nearly three months, and the agency announced plans last week to try to restore contact with it. Jeff Foust reports that the overall Mars exploration program at NASA is facing challenges as well.
Tuesday, September 4, 2018
http://thespacereview.com/article/3563/1
Astronomers use Earth’s history as guide to spot vegetation on new worlds
By Blaine Friedlander
September 24, 2018
By looking at Earth’s full natural history and evolution, astronomers may have found a template for vegetation fingerprints – borrowing from epochs of changing flora – to determine the age of habitable exoplanets.
“Our models show that Earth’s vegetation reflectance signature increases with coverage of our planet’s surface, but also with the age of our planet,” said co-author Jack O’Malley-James, research associate in astronomy at Cornell’s Carl Sagan Institute. The research, “The Vegetation Red Edge Biosignature Through Time on Earth and Exoplanets,” published online Sept. 12 in Astrobiology Journal.
Full article here:
http://news.cornell.edu/stories/2018/09/astronomers-use-earths-history-guide-spot-vegetation-new-worlds
To quote:
“We use Earth’s history as a key for finding life in the universe,” said co-author Lisa Kaltenegger, associate professor of astronomy and director of the Carl Sagan Institute. “Our work shows that as plants evolved on Earth, the vegetation signal that reveals their presence became stronger, making older exoplanets really interesting places to look for vegetation.”
Exoplanets may be parched, arid with clear skies and endless cacti forests, or hot jungle worlds covered in tropical forests, said Kaltenegger, “Over interstellar distances, these places might be the best targets to spot vegetation.”